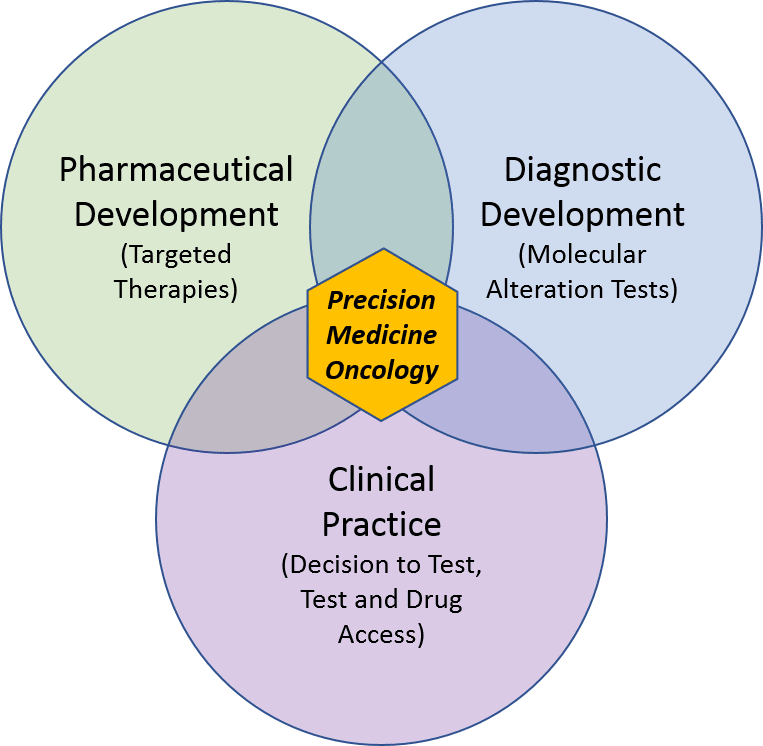
Over the past few decades in oncology, targeted therapies—and the tests that identify which tumors will be sensitive to them—have significantly advanced the standard of care in several key settings. However, precision medicine will not achieve its full potential in oncology without broader success in pharmaceutical and diagnostic development, and better integration of both into clinical practice.
In Part I of our 3-part series on precision medicine oncology, we reviewed the scientific basis of molecular alterations and the technology of the tests that detect them. Here, we explore how precision medicine is used in the clinic, as well as the key challenges associated with it. In Part III we will focus on the future landscape and the requirements for both diagnostics and pharma to achieve broader success.
Key terms:
- Diagnostic: a test for molecular alterations that indicates or confirms a disease or specific molecular alteration
- Molecular alteration: any change in a protein or its precursors that impedes its normal biological function; This includes both mutations that change the function of a protein, as well as significant increases or decreases in the amount of the protein.
- Precision medicine: collecting information about molecular alterations and incorporating it into treatment decision-making
- Predictive vs Prognostic test: Predictive tests indicate whether a particular treatment is more or less likely to work for the tumor, whereas prognostic tests provide information about the likely course of disease (treatment agnostic).
Current Application of Precision Medicine in Oncology
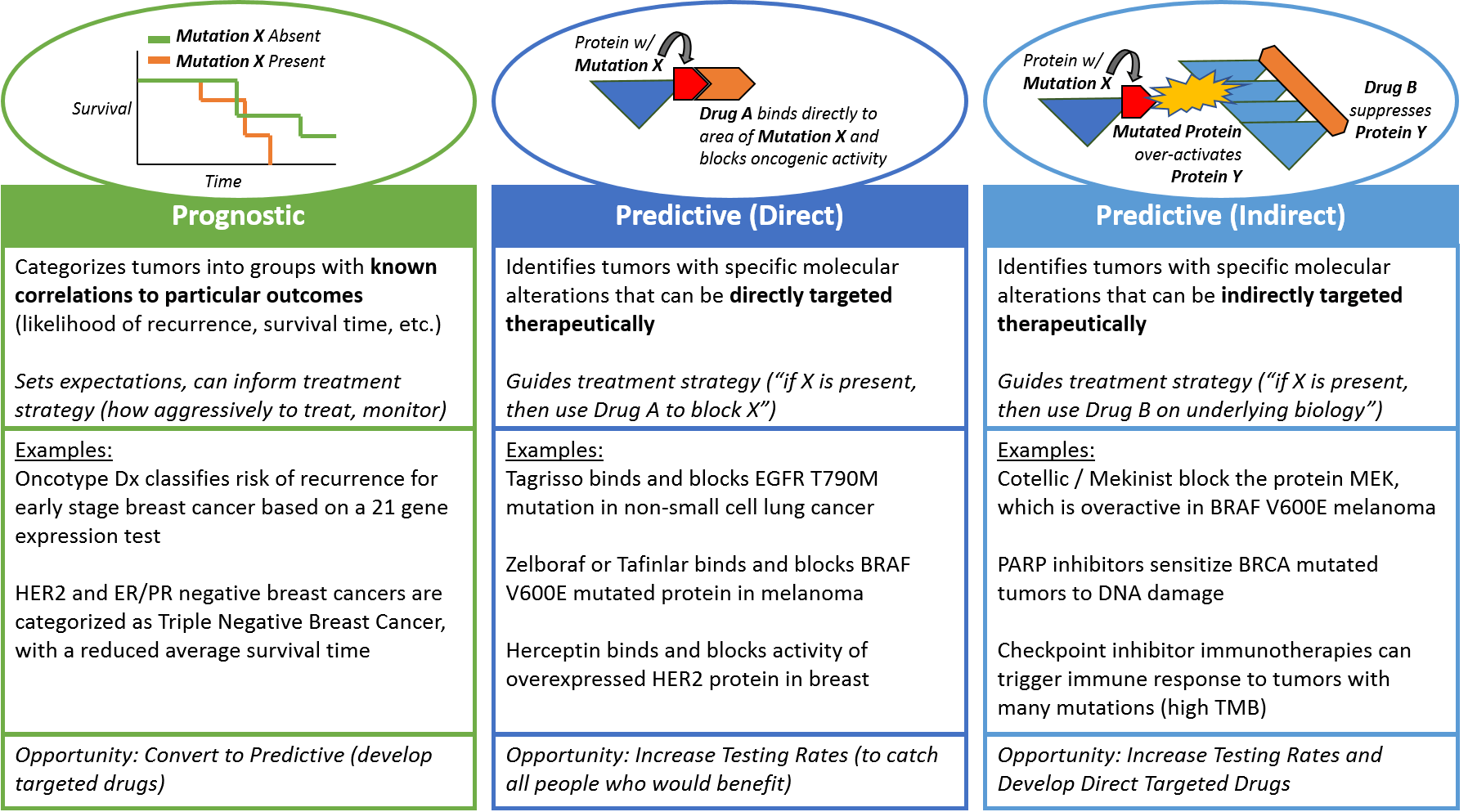
Currently, precision medicine is used in oncology in three distinct manners: prognostic, predictive (direct), and predictive (indirect). While all three types of test focus on classifying the underlying biology of the tumor, the clinical utility of each is somewhat distinct.
Simply put, prognostic tests provide additional information about disease characteristics beyond what pathology tests can show, whereas predictive tests indicate sensitivity to a particular intervention – whether it will have a higher (or lower) likelihood of effectiveness for this particular tumor.
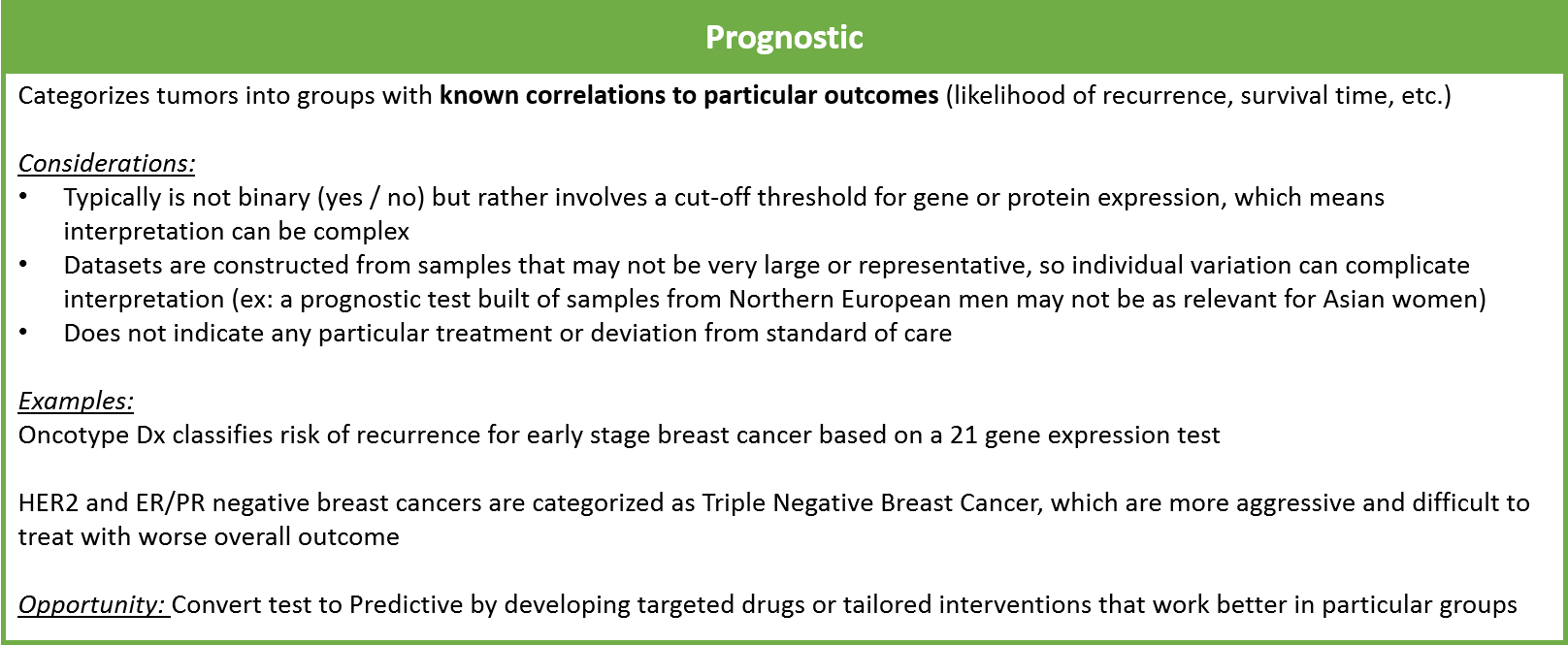
Prognostic tests provide information about the likely course of disease under the standard treatment paradigm. For example, the Oncotype Dx test uses a panel of 21 mRNA expression tests to categorize early stage breast tumors into high, intermediate, or low risk of recurrence after standard treatment. In later-stage breast cancers, the absence of both HER2 and ER/PR protein overexpression is used to categorize tumors as “Triple Negative”, with a worse prognosis.
Prognostic tests do not suggest a specific treatment but can help guide overall treatment strategy and set expectations for the patient and loved ones. Ideally, prognostic tests will evolve over time to become predictive as new treatments are developed.
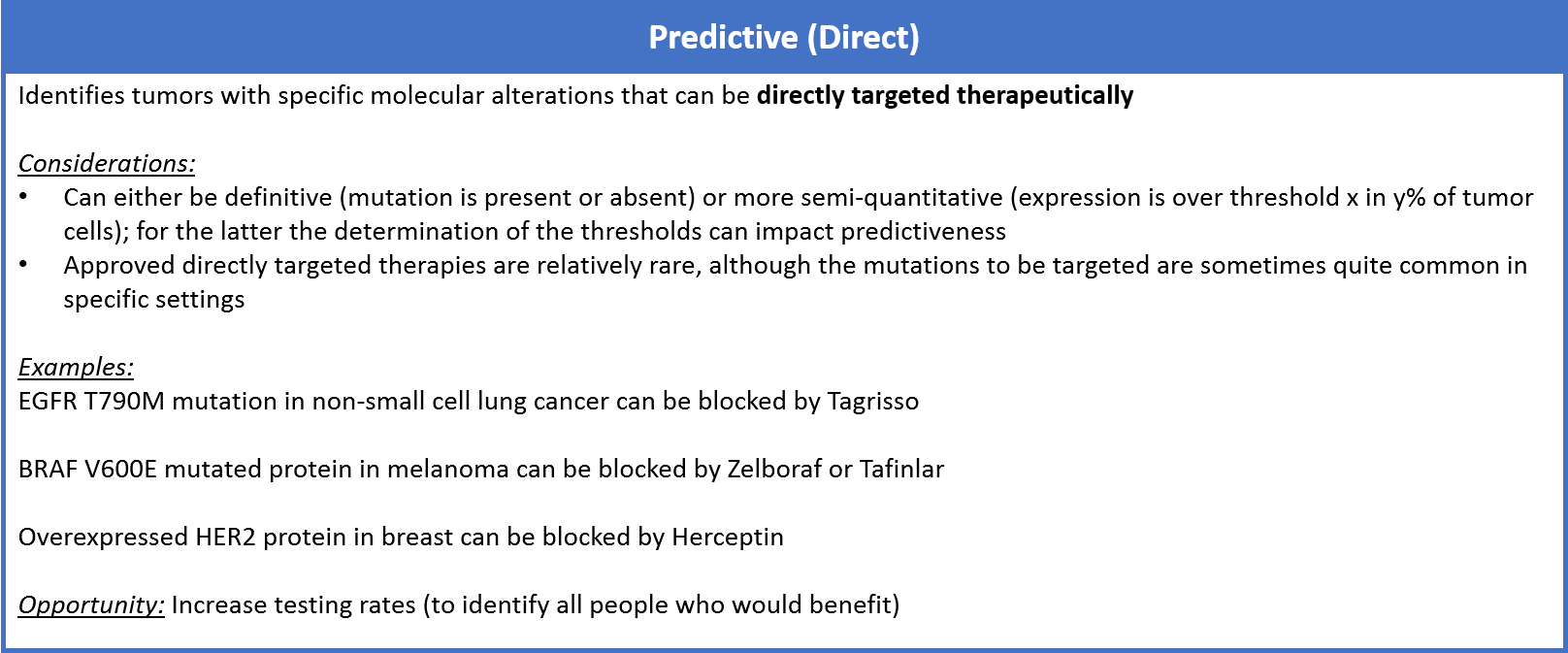
Predictive tests also categorize tumors based on patterns of molecular alterations, but unlike prognostic tests they suggest specific treatment approaches to address the underlying biology. Predictive tests indicate treatments that can be classified as direct or indirect.
Direct targeted drugs are those that directly interact with and negate the harm of mutated or mis-regulated proteins. For example, the drug Tagrisso binds specifically to the T790M mutation on the protein EGFR, which arises as a mechanism of resistance to first-line EGFR inhibitors in non-small cell lung cancer. Other examples include the BRAF inhibitor drugs, Zelboraf or Tafinlar, used in melanoma to specifically target the BRAF V600E mutation. Another common example is the HER2 targeted drugs (like Herceptin) which are used to block HER2 overexpression if it is identified at the gene or protein level in breast cancer. Typically, direct targeted drug efficacy is very well-correlated with the predictive marker.
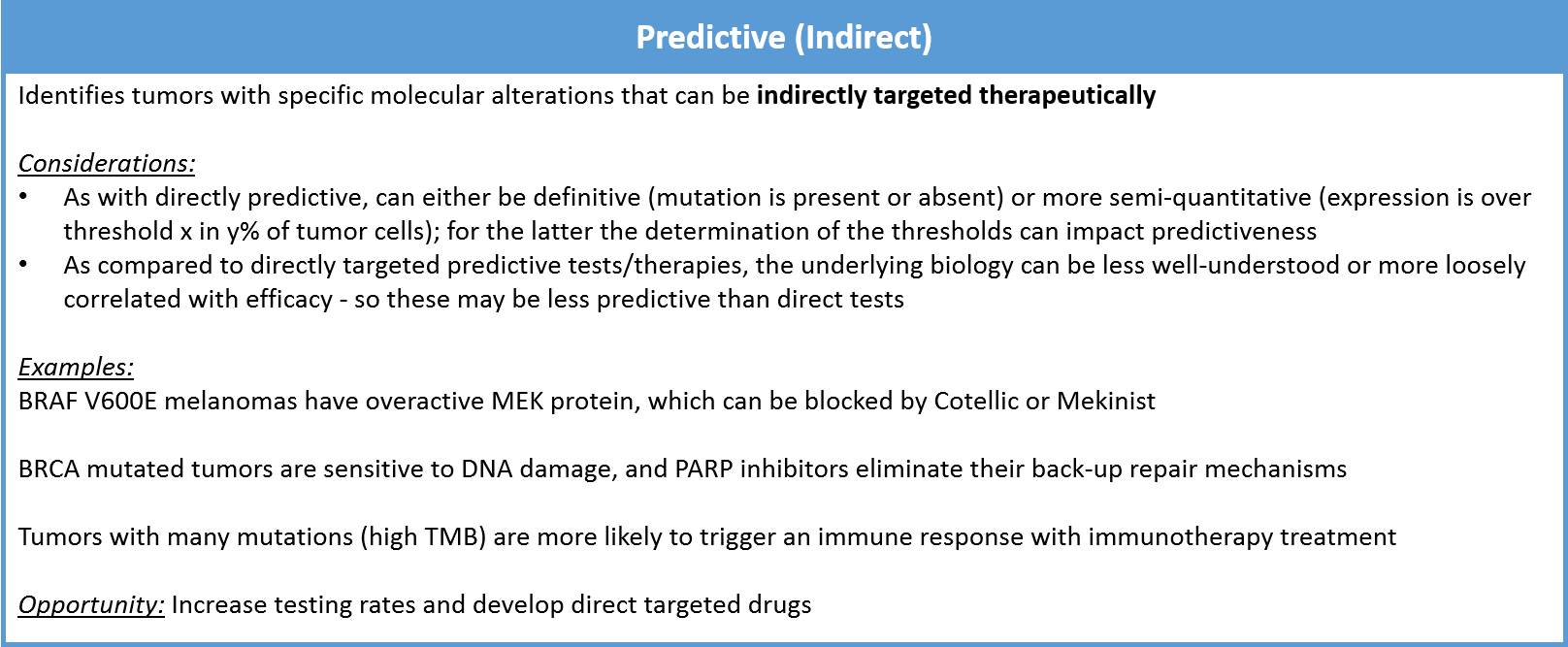
An indirect predictive test still indicates whether a particular therapy is more (or less) likely to work, but it does not measure the target directly. Instead, it surveys a separate but strongly-correlated molecular alteration that is an indirect predictor of response. This can be another protein that operates nearby in the same oncogenic pathway. For example, the MEK inhibitor drugs Cotellic or Mekinist are often used in BRAF V600E positive melanoma. They work indirectly by suppressing the overactivity of MEK, which occurs when BRAF has the V600E mutation.
Indirectly predictive tests can also illuminate the underlying biology rather than a single protein misregulation. For example, tumors that have a BRCA mutation are unable to repair certain types of DNA damage. In this setting, drugs called PARP inhibitors can be used to block their back-up repair mechanisms. This can dramatically improve the efficacy of DNA-damaging systemic treatments like radiation or chemotherapy.
The reason to differentiate direct vs indirect in predictive tests is that a direct test is typically easier to interpret and has a higher correlation with efficacy. When either designing or choosing between tests, a direct predictive marker is preferable, if feasible.
Checkpoint immunotherapies are an interesting case of a therapy that can be indicated by either a direct or indirect predictive test, depending on the molecular alteration identified. If a tumor has PDL1 overexpression, then the PD1/PDL1-targeted checkpoint inhibitors (ex: Keytruda, Opdivo, Tecentriq) can directly suppress the inappropriate activity of the PDL1 checkpoint.
If a tumor has high tumor mutation burden (TMB), this indirect marker indicates that there are numerous mutations throughout the genome that make it more likely for the immune system to recognize and attack the cancer. In such a case, checkpoint inhibitors are more likely to kickstart the process of immune activation. PDL1 expression and TMB therefore illuminate different underlying biologies and are independent predictors of response to checkpoint immunotherapies.
In general, directly predictive tests and their associated targeted therapies are the gold standard (since the drugs act directly on the alteration measured by the test). Indirect predictive tests can also be quite useful, but only if there is a high enough correlation between test result and drug efficacy.
Prognostic tests have relatively limited utility in clinical decision-making. However, they can be quite helpful to patients trying to appropriately manage their expectations for the course of their disease.
Integration of Precision Medicine into Clinical Oncology Practice
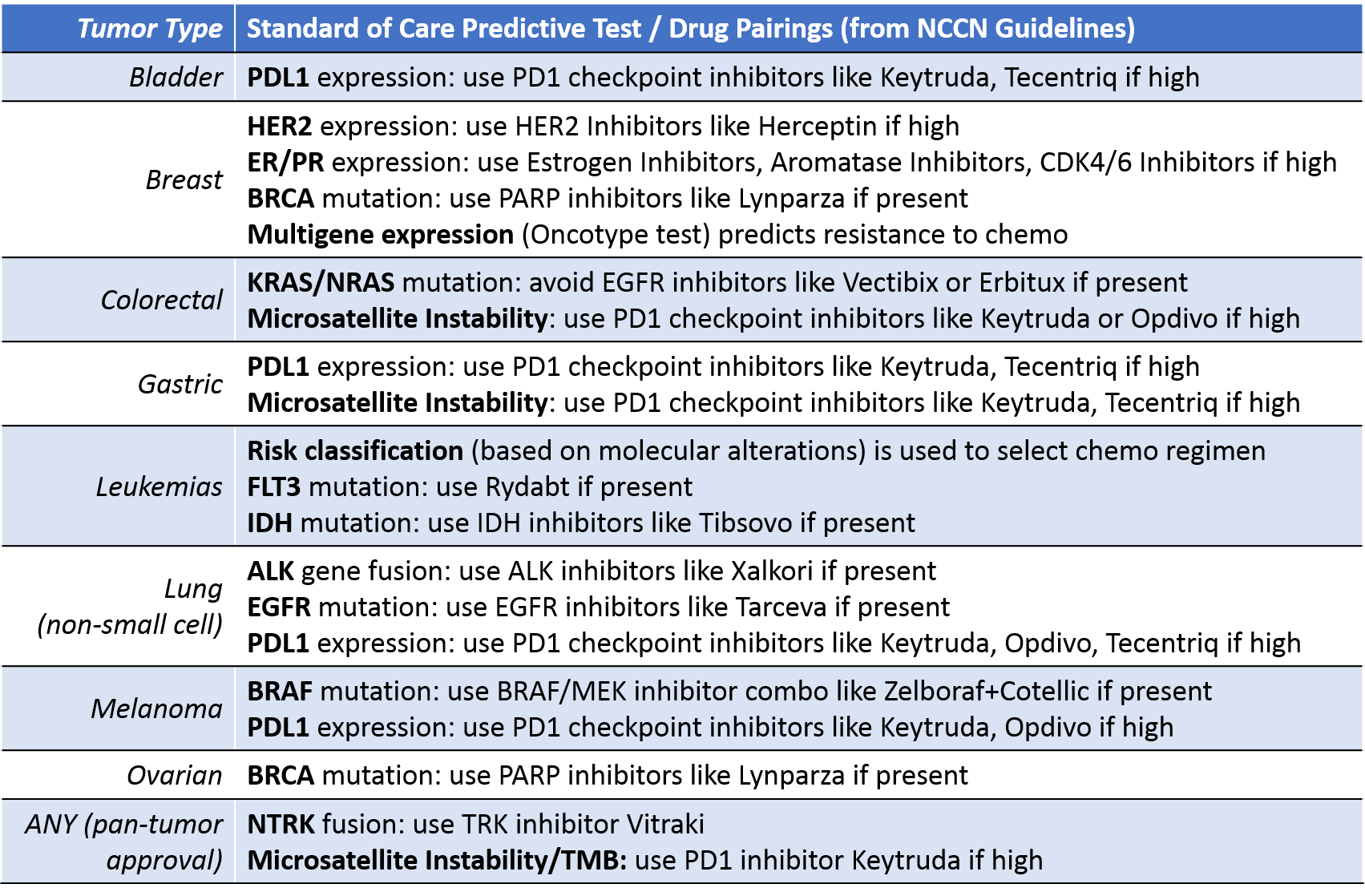
Currently, precision medicine is fully integrated into the standard of care for cancer in only a few settings, those in which:
- Particular molecular alterations are relatively common, and
- Predictive test/drug pairings can successfully target that alteration
There are also two recent examples of drugs that have achieved pan-tumor indications based purely on predictive testing:
- VITRAKVI for any solid tumor with NTRK gene fusion
- Keytruda for any solid tumor with high TMB / genomic instability (Microsatellite Instability High or MMR deficient).
A wide breadth of tumor types have at least some element of precision medicine integrated into the standard of care. Within each tumor type, however, there is high variability in how many patients receive testing. This is can be driven by:
- Narrow indications (for example testing is only done in certain lines of therapy or for patients who have already received other specific treatments)
- Gaps in physician awareness
- Limited patient access to tests even within approved indications
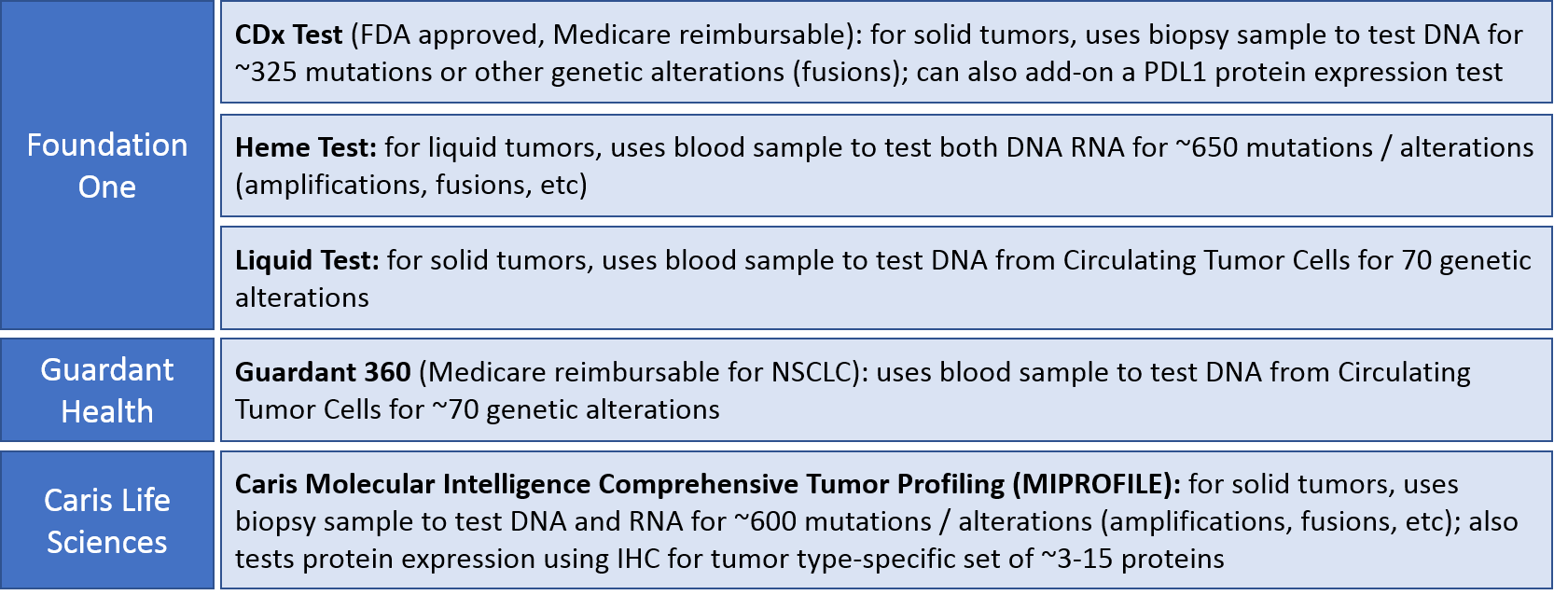
In addition to the very targeted tests described above, broader panel tests are emerging that typically include numerous genetic alterations (DNA testing). Some can also include a few common protein expression alterations assessed by IHC. These tests identify molecular alterations that can indicate approved, off-label, or experimental (clinical trial) treatments.
These broader panel tests are not yet officially part of the standard of care. However, academic centers use them rather often, and some have their own in-house versions (ex: UCSF500 gene panel). They’re also sometimes used in rare cancers or later lines of therapy where there is the hope of identifying a rare but actionable alteration. Finally, these tests are sometimes requested by patients interested in proactively seeking treatment options beyond the standard of care.
For these panel tests, the interpretation is almost always simple 1:1 mapping of regimens to single alterations (use – or don’t use – Drug X because of Mutation Y). In the cases where multiple molecular alterations are discovered, the clinician receives a list of alterations and drugs or clinical trials to consider. The clinician is then responsible for interpreting which results are most meaningful and prioritizing treatment options accordingly. Needless to say, this can be quite challenging.
The broader panel tests also include many alterations that are not fully predictive. They may provide prognostic information, hint at underlying biology that could be addressed therapeutically, or have targeted therapies in clinical trials.
Key Challenges in Advancing Precision Medicine
To advance precision medicine, we must accelerate pharmaceutical development, diagnostic development, and integration into clinical practice. Simply put, we need more and better targeted drugs, more predictive tests to guide their usage, and better uptake of both the tests and drugs in clinical practice.
The underlying barriers to achieving all the goals above relate to scientific and technical issues, clinical development challenges, and commercial issues. Some may be solvable given enough resources and time; others (like developing a targeted drug for a particular alteration) may be ultimately prove intractable.
One critical unmet need is better pharmaceutical approaches to target known drivers of malignancy. For example, overactivity of the protein MET, driven by gene amplifications and / or protein overexpression, is a common molecular alteration in many tumors, particularly lung, colorectal, and liver. None of the numerous attempts to develop MET inhibitors in clinic have yet succeeded. This has not been a failure of drug design per se (the drugs do inhibit MET protein as intended). Rather, it indicates some gap in scientific understanding – either MET protein overactivity is not truly a primary driver of malignancy, or perhaps clinical efficacy will require more complex approaches. These could include combinatorial or sequential treatments to address other parts of the MET pathway, a better predictive test to identify likely responders, etc.
To achieve clinical efficacy with regards to this pathway—and others where drug development has so far underperformed—we need a deeper understanding of tumor biology that will clarify a better approach. Perhaps that would be a different type of inhibitor, a combination treatment strategy, or a pattern of background genetics that creates susceptibility, etc.
In parallel, diagnostic development has struggled to support some drugs and classes of drugs where we either have only indirect predictive tests, or none. The immunotherapies are a notable example. Both the checkpoint inhibitors as well as the promising but less developed vaccine, virus, and cell-based therapy approaches show a distinctive “long tail” response curve where a small percentage of recipients have an exceptionally durable (even curative) response to treatments. However, the majority suffer the side effects of these therapies but receive no real benefit.
While tumor mutation burden and PDL1 expression are considered predictive for response to checkpoint inhibitors, they are not well-correlated enough for many clinicians to risk undertreating. Instead, they treat everyone, knowing that there can still be some responders in the “low” test result category.
There are also some therapies which are targeted in the sense that they directly block a protein, but alterations of that protein do not appear to be predictive. Avastin and the other VEGF inhibitors are a notable example where diagnostic tests have never been successfully developed. Better predictive tests would result in a more favorable benefit/risk ratio for these drugs, improving outcomes for more patients as well as cost savings where expensive drugs can be safely avoided.
The underlying barriers to diagnostic development are typically associated with test design choices. If a test is intended to complement an existing drug, then the molecular alteration is likely already known. However, diagnostic test designers may still need to choose which type of assay (DNA, mRNA, protein) will provide the best balance between accuracy and feasibility. For example, protein tests inherently require more sample and more qualitative interpretation, so DNA-based tests may be preferred when possible.
For the broader, panel-based DNA tests that are not aligned to a particular drug, there is the question of which genetic alterations to include. There are ~20,000 genes in the human genome, each of which can exist in numerous mutated or otherwise forms. Panel tests typically restrict themselves to a list of a few hundred or dozens of genetic alterations that are either prognostic or predictive / targetable.
The challenge is that new information is constantly coming out, and updates to tests—and their interpretation—must be carefully considered. Updating too frequently can confuse consumers. Updating too infrequently can let the tests get outdated.
The other somewhat open question is how slightly modifying existing tests will impact their FDA approval and reimbursement status. Will they need to re-demonstrate clinical utility with each new version?
As clinicians attempt to integrate drugs and tests into their practice, they face barriers of awareness, access, and interpretation. Even in cases where testing is part of the standard of care and has been for some time (for example EGFR testing in lung), the testing rate is still not 100%. Ongoing consumer education of both physicians and patients will continue to be required to maximize testing rates.
Given the complexity of precision medicine testing technology, clinicians are challenged to decide whether to test, and which test to use. There are some clear-cut contexts where a companion diagnostic is labeled for use to select a specific drug. Examples include the Cobas blood-based DNA sequencing test that detects the EGFR T790M mutation (which the drug Tagrisso can target) or the various PDL1 tests marketed for use with checkpoint inhibitor immunotherapies. However, in the majority of treatment settings, the standard of care guidelines do not recommend specific precision medicine tests, so it’s at the discretion of the clinician.
Even if the clinician and / or the patient is aware of and wants to test, there may be access barriers associated with feasibility of obtaining an appropriate sample. For example, a progression may be identified on an MRI but a biopsy is not done because the pathology is already inferred.
The more common access barrier is reimbursement and cost, particularly for those tests that have not provided clinical utility nor obtained FDA approval. Many testing companies provide financial support to patients as needed, but this is not sustainable for growth. A related barrier can arise when a test identifies an actionable alteration, but the drug is not approved in that indication. This can create both logistical and financial hurdles to access the drug off-label.
Interpretation is a very challenging barrier in certain settings. Specifically, it’s challenging in settings where testing is done but the results are either too vague (nothing much actionable) or too complicated (multiple actionable targets identified).
In the case of the former, a clinician can always revert to the standard of care, but there may be a missed opportunity of an actionable alteration wasn’t tested for or that wasn’t recognized as potentially actionable in the read-out. For example, a set of several alterations within the same pathway might indicate that the pathway should be targeted, but a clinician would have to make that inference on their own because test reports typically only provide 1:1 alteration:drug information.
In the latter case, if more than one obviously actionable alterations are identified, the test reports still only provide the 1:1 information, i.e., a list of each alteration identified along with appropriate drugs or clinical trials for each. The reports do not, however, provide any relative prioritization that would support a clinician’s decision between two or more options.
Conclusion and Future Directions:
Maximizing the potential of precision medicine will require co-evolution of diagnostic and pharma to boost the utility of both, as well as a higher order understanding of how to integrate test results into clinical practice. In our next installment, we will explore how diagnostic and pharma companies can lead the advancement of precision medicine.